Fluorescent In Situ Hybridization: (FISH)
What Is FISH?
FISH is a molecular technique that is often used to identify and enumerate specific microbial groups. This technique can be used to determine, with the presence or absence of a fluorescent signal, whether specific genetic elements exist in a sample. This can be useful for determining if microbes have a particular gene present and/or if that gene is being expressed under a given set of conditions. Fluorescent probes are designed to attach to specific genetic regions of microbes that will differentiate them from other groups. When these probes are applied a fluorescent microscope can be used to detect the presence or absence of individual microbial groups. Another sister technique, called Flow-Cytometric Analysis (FCM), can also be carried out when fluorescent tags are applied to microbial populations. Fluorescent signal is used to count or sort individual genotypes out of groups of cells (see more on FCM).
How Does FISH Work?
The FISH technique is dependent upon hybridizing a probe with a fluorescent tag, complementary in sequence, to a short section of DNA on a target gene. The tag and probe are applied to a sample of interest under conditions that allow for the probe to attach itself to the complementary sequence in the specimen if it is present. After the specimen has been treated, excess fluorophore is washed away and the sample can be visualized under a fluorescent microscope. By quantifying the amount of fluorescence with the scope it can be determined if the type of cell the probe was designed for is present, and if so, how much of it is present in a sample. Transcripts (mRNA) in microbes can also be targeted to detect if a specific gene is being expressed under the given conditions.
Applications:
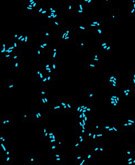
Fluorescent probes of various colors can be used at the same time for varying targets at the same time to determine which portion of a population different individuals make up. For instance, in the image shown here, a red archaeal specific probe has been applied as well as green bacterial probe. These images can be compared with those taken with a different filter and stain such as DAPI that binds DNA nonspecifically. The images with the blue DAPI stain show the size of the combined microbial populations and when compared with the differentially stained images of the same population can give researchers an idea of what proportion of the whole each of the different domains (bacteria and archaea) are responsible for.
Related Links
- FISH - use of the technique in looking at the evolution of complex systems
- Nikon Microscopy U - hardware and software implications in the research laboratory